Cancer AI investment continues to pour into models and raw data-foundation architectures, multimodal systems, and ever-larger training datasets. Yet despite the surge of AI-enabled medical devices, the gap between research performance and clinical deployment remains persistent. Models that achieve impressive accuracy in controlled environments often fail in community hospitals. Authorized systems sit unused. Cross-border data collaboration remains operationally fragile.
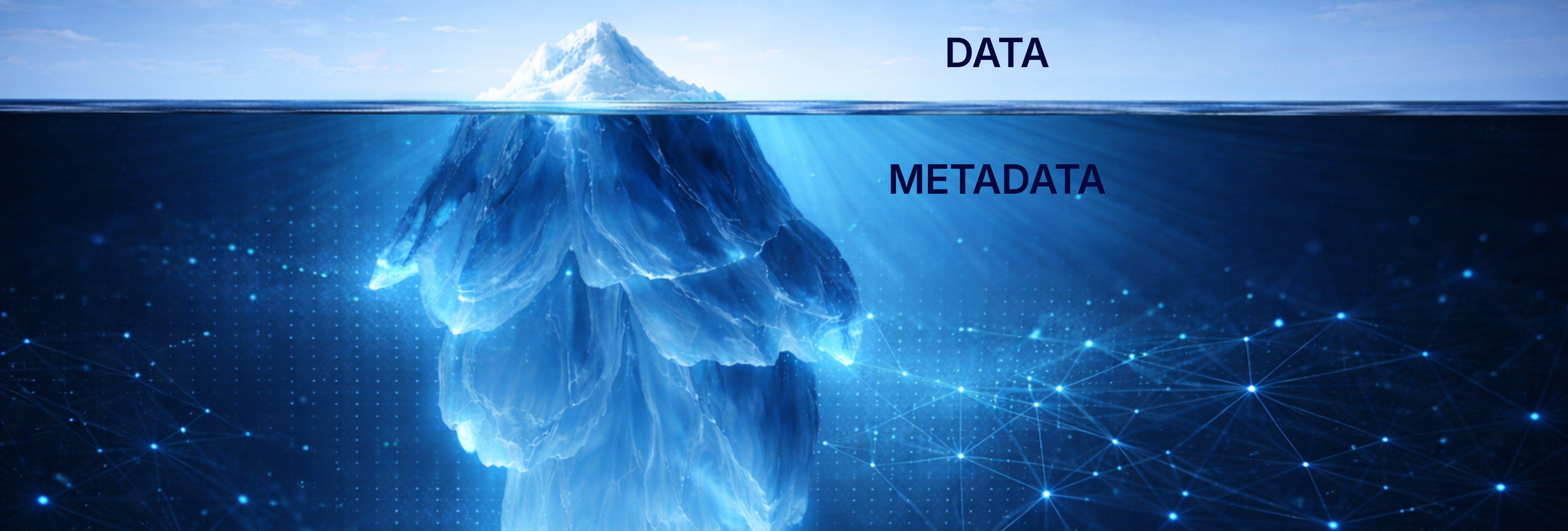
The constraint is no longer algorithmic sophistication. It is the data infrastructure beneath the algorithms, specifically, metadata that determines whether cancer AI scales, stalls, or produces sustainable return on investment.
What Metadata Really Means in Healthcare AI
Metadata is often described as "data about data." In cancer AI, that definition is inadequate. Metadata is the biography of a data point, the structured context that transforms raw pixels, clinical text, or molecular measurements into clinically interpretable evidence.
In oncology, that context spans multiple layers. Patient and clinical metadata includes demographics, diagnosis, staging, treatment history, and outcomes. Specimen and laboratory metadata covers tissue origin, fixation method, staining protocol, and processing parameters. Diagnostic metadata captures pathologist assessments, tumor grading, biomarker quantification, and synoptic report elements. Technical and AI-specific metadata documents scanner configuration, acquisition settings, annotation provenance, preprocessing pipelines, and model versioning.
Cutting across all layers is the patient journey, including follow-up intervals, treatments, censoring indicators. Since cancer unfolds over months to decades, metadata enriches the understanding of clinical outcomes and their determinants.
The challenge is structural fragmentation. Laboratory information systems, electronic health records, scanner software, paperwork each store contextual fragments in different formats, vocabularies, and standards. Without structured, validated metadata, AI systems are biased and inaccurate.
The Hidden Cost of Data Debt
For executive leadership and investors, the absence of metadata infrastructure is not just a compliance issue; it is financial liability.
Healthcare organizations often prioritize acquiring raw data, petabytes of whole slide images or genomic sequences, assuming value lies in scale. Yet raw data without validated metadata is a depreciating asset. When a regulator requests dataset lineage, when a pharma partner demands cohort reconstruction, or when a model requires retraining, retrospective reconstruction begins.
Highly trained pathologists and data scientists become data archaeologists.
This accumulated burden is data debt. It compounds silently until a strategic moment exposes it during due diligence, regulatory review, or clinical scale-up. Retrospective curation is slow, expensive, and error-prone.
Automated metadata validation at ingestion converts data into a liquid asset. Cohorts become queryable. Provenance becomes traceable. The question shifts from "Can we reconstruct this dataset?" to "Which validated population should we analyze today?"
In the current market, valuation increasingly reflects dataset defensibility, rather than dataset volume. Metadata maturity directly affects enterprise value.
When Metadata Breaks, Generalization Breaks
The central risk in cancer AI is not architectural weakness, but hidden confounding that limits generalization. A systematic review of AI in digital pathology reported accuracy reductions of 6% to 23% in the absence of image normalization and demonstrated that models trained on pathology slides can predict scanner type, tissue thickness, and processing laboratory, none of which are biologically meaningful [1]. Reported performance is further inflated by data leakage, the illicit sharing of information between training and test data, a pervasive problem across machine learning in the biological sciences [2].
The risk emerges when acquisition-specific features correlate with diagnostic labels in the training data. Technical variables become unintended shortcuts. Internal validation appears strong. External performance deteriorates.
The problem runs deeper than individual confounders. A majority of published pathology AI studies fail to report fixation and preparation details [1], making external validation unreliable. Performance gaps become impossible to diagnose: are they biological, technical, or procedural? Survival models depend on censoring indicators and follow-up completeness, yet informative censoring where patients leave follow-up for reasons correlated with their outcomes introduces systematic bias that few studies quantify [3].
This is not a documentation problem; rather it is an engineering problem. Manual metadata curation cannot scale across the volume and heterogeneity of modern pathology data. What is needed is automated extraction from diverse sources, vocabulary normalization against standard terminologies, and systematic quality assurance at the point of ingestion, rather than retrospective cleanup.
Accelerating Pharma R&D: Metadata as a Trial Engine
For pharmaceutical organizations, metadata directly impacts time-to-market.
Precision oncology trials increasingly depend on granular inclusion criteria: molecular subtype, biomarker expression thresholds, prior immunotherapy exposure, staging definitions, and longitudinal outcomes. Identifying such cohorts across distributed systems is slow when data remains unstructured.
A harmonized metadata layer transforms feasibility analysis into a computational process. Instead of manual chart review, pharma teams can query validated metadata across networks to determine how many eligible patients exist, and where.
Companion diagnostics depend on this same infrastructure. Demonstrating that a biomarker correlates with clinical outcome requires linking slide-level data, molecular findings, and longitudinal survival data in a reproducible manner. Without structured metadata bridging these domains, regulatory-grade evidence becomes fragile.
Real-world evidence generation further depends on harmonized outcome definitions across jurisdictions. When "progression-free survival" varies by institution or country, pooled analysis loses validity.
Metadata is therefore not an administrative layer in drug development. It is the operational substrate of precision medicine.
Multimodal AI: Integration Requires Alignment
The next frontier in oncology is multimodal AI, integrating pathology, radiology, genomics, proteomics, and longitudinal clinical history.
Multimodal systems do not fail at the modeling layer. They fail at the alignment layer.
A slide image cannot be meaningfully fused with a genomic sequence unless specimen lineage and temporal relationships are encoded. Was the biopsy obtained before therapy initiation? Does the sequencing derive from the same tissue block as the digitized slide? Are staging definitions harmonized across institutions?
Metadata encodes these dependencies and defines comparability. Without it, multimodal AI remains an academic demonstration. With it, integration becomes clinically reliable and regulatory-ready.
Operational Traceability: The Audit Beyond Regulation
European regulators are making metadata an explicit requirement for high-risk medical AI. The EU AI Act (Regulation 2024/1689), with enforcement beginning August 2026 for standalone healthcare AI, mandates data governance under Article 10: training datasets must be relevant, representative, and free of errors, with documented collection processes, preparation operations, and statistical properties regarding target populations [4]. MDCG 2025-6 establishes that AI embedded in medical devices must satisfy both MDR and AI Act requirements simultaneously, with data traceability treated analogously to material traceability in traditional devices [5]. The European Health Data Space (Regulation 2025/327), which entered into force in March 2025, mandates Health Data Access Bodies by 2027 and cross-border health data exchange by 2029, creating an operational framework where metadata standardization is legally required [6]. FDA lifecycle guidance provides complementary international alignment [7].
However, traceability is not solely a regulatory requirement; it is an operational necessity.
Structured metadata enables rapid attribution of variance and interruptions to specific operational variables. In high-throughput oncology environments, this capability determines whether AI systems remain trusted clinical tools or fragile technical experiments.
Despite these requirements, the evidence base remains thin. A cross-sectional study of 691 FDA-cleared AI devices found that 47% of summaries lack study design descriptions and 95% fail to report demographic representation in training data [8].
Field-level provenance, which traces every data point from raw source through normalization, validation, and into model training, is not optional documentation. It is the operationalized evidence that regulatory compliance demands. Compliance infrastructure must be built into the data pipeline, not bolted on afterward.
Equity and Global Deployment
Cancer AI must serve diverse populations, not just the institutions where models are built. Computational pathology models display marked performance disparities across demographic groups, with AUC gaps of up to 16% for cancer subtyping, and disparities identified in 29.3% of diagnostic tasks across 20 cancer types [9, 10]. Fairness-aware training strategies are emerging, yet they depend fundamentally on demographic and acquisition metadata.
The root of this problem is data concentration: a systematic review of clinical text datasets found that 73% originate from the Americas and Europe, regions representing only 22% of the global population [11]. Without demographic and geographic metadata, these imbalances remain invisible.
The European Health Data Space mandates cross-border data exchange, and initiatives like EUCAIM are building the imaging infrastructure to support it [12]. But cross-border AI requires more than shared repositories; it requires harmonized metadata across institutions with different coding systems, languages, and laboratory workflows.
Equitable AI requires metadata that makes diversity measurable across demographic, geographic, and institutional dimensions. This demands automated harmonization that normalizes vocabulary across sources while preserving the provenance of every transformation. Fairness cannot be an afterthought appended to a model card; it must be embedded in the data infrastructure.
From Algorithms to Infrastructure Thinking
Cancer AI is entering its infrastructure phase. The bottleneck has moved upstream: fragmented data across laboratory systems, clinical records, scanner formats, and genomic pipelines.
The limiting factor is no longer model novelty. It is system maturity, defined by data governance, ontology management, validation discipline, and lifecycle traceability.
Pharmaceutical acceleration, regulatory readiness, multimodal integration, operational resilience, and equitable deployment are not independent objectives. They are consequences of structured metadata governance.
At PAICON, we build AI systems with this premise from the outset: models evolve, but infrastructure determines durability. Automated, AI-assisted metadata pipelines that extract, validate, normalize, and harmonize heterogeneous oncology data under human oversight transform siloed information into structured, research-ready assets that meet regulatory requirements and support equitable AI deployment.
The next generation of cancer AI will not be defined by algorithmic novelty or who trains the largest model. It will be defined by the strength of the infrastructure beneath it that survives real clinical complexity. And metadata is that infrastructure layer.
Build on Infrastructure That Scales
Access harmonized, metadata-structured multinational cancer datasets enabling scalable AI validation, benchmarking, and regulatory-grade deployment across real-world clinical environments.
References
1. McGenity C, Clarke EL, Jennings C, et al. Artificial intelligence in digital pathology: a systematic review and meta-analysis of diagnostic test accuracy. npj Digit Med. 2024;7:114. DOI: [10.1038/s41746-024-01106-8](https://doi.org/10.1038/s41746-024-01106-8)
2. Bernett J, Blumenthal DB, Grimm DG, et al. Guiding questions to avoid data leakage in biological machine learning applications. Nat Methods. 2024;21(8):1444–1453. DOI: [10.1038/s41592-024-02362-y](https://doi.org/10.1038/s41592-024-02362-y)
3. Templeton AJ, Amir E, Tannock IF. Informative censoring — a neglected cause of bias in oncology trials. Nat Rev Clin Oncol. 2020;17:327–328. DOI: [10.1038/s41571-020-0368-0](https://doi.org/10.1038/s41571-020-0368-0)
4. European Parliament and Council. Regulation (EU) 2024/1689 laying down harmonised rules on artificial intelligence (AI Act). Off J Eur Union. 2024;L 2024/1689.
5. Medical Device Coordination Group. AIB 2025-1, MDCG 2025-6: Interplay between MDR/IVDR and the AI Act (AIA). European Commission; 2025.
6. European Parliament and Council. Regulation (EU) 2025/327 on the European Health Data Space (EHDS). Off J Eur Union. 2025;L 2025/327.
7. U.S. Food and Drug Administration. Artificial Intelligence-Enabled Device Software Functions: Lifecycle Management and Marketing Submission Recommendations. Draft Guidance. FDA; 2025.
8. Lin JC, et al. Benefit-risk reporting for FDA-cleared artificial intelligence-enabled medical devices. JAMA Health Forum. 2025;6(9):e253351. DOI: [10.1001/jamahealthforum.2025.3351](https://doi.org/10.1001/jamahealthforum.2025.3351)
9. Vaidya AJ, Chen RJ, Williamson DFK, et al. Demographic bias in misdiagnosis by computational pathology models. Nat Med. 2024;30:1174–1190. DOI: [10.1038/s41591-024-02885-z](https://doi.org/10.1038/s41591-024-02885-z)
10. Lin SY, Tsai PC, Su FY, et al. Contrastive learning enhances fairness in pathology artificial intelligence systems. Cell Rep Med. 2025;6(12):102527. DOI: [10.1016/j.xcrm.2025.102527](https://doi.org/10.1016/j.xcrm.2025.102527)
11. Wu J, Liu X, Li M, et al. Clinical text datasets for medical artificial intelligence and large language models — a systematic review. NEJM AI. 2024;1(6):AIra2400012. DOI: [10.1056/AIra2400012](https://doi.org/10.1056/AIra2400012)
12. EUCAIM Consortium. European Cancer Imaging Initiative: Infrastructure and Standards Report. European Commission; 2024.